Fionn Murtagh's
Multivariate
Data Analysis
Software and Resources
Page
Contents
- Hierarchical Clustering Software in R/S-Plus
- MDA-J: Multivariate Data Analysis - Java
- Multivariate Data Analysis Software as Standalone Java Applications
- Gaussian Mixture Modeling with Bayes Factors in C
- Wavelet Transform on a Hierarchy or Dendrogram
- Multivariate Data Analysis Software in Fortran (and C)
- Resources and Links, including: JP Benzécri's Pascal Code
for Correspondence Analysis; Multidimensional Scaling;
Point Pattern Matching; book F. Murtagh, Multidimensional
Clustering Algorithms, Physica-Verlag, 1985;
Reading FITS Files in R.
- Software accompanying
the book:
Correspondence Analysis and Data Coding with R and Java
Legal notice: all software here can be freely used and incorporated
into any system, for
any purpose. It is required that the origin and authorship of any code
taken from here is acknowledged in code and other documentation.
1. Hierarchical Clustering Software in R/S-Plus
What is unique here: (1) All hierarchical clustering programs achieve
the optimal O(n2) computational bound using the nearest neighbors
chain algorithm. (2) The "stored dissimilarity" algorithm is used,
implying O(n2) storage (the "stored data" algorithm is an
alternative, with O(n) storage, but greater absolute computational
requirement).
(3) For hierarchical clustering, both
native R code and linked C code (crucial for efficiency, with large
data sets);
implementations are identical.
(4) Weighting of cases/observations (rows) supported. (5) Easy and
straightforward linkage with correspondence analysis to normalize (by
"Euclideanizing") the data input to the hierarchical clustering. (6)
The Ward minimum variance criterion is used in the software here. (See note
below.)
-
"Readme": Background details, and
examples of use on Intel Windows and Unix (including Mac OS X) systems.
- All
functions in one (standalone) R/S-Plus file
(call in R as function hierclust).
- Hierarchical clustering in C, used in R environment.
- Normalizing input data
- Correspondence analysis area
(including lots of data sets).
- The hierarchical clustering is identical to what has been listed in
this section.
- The correspondence analysis is used in R to "Euclideanize"
the data (e.g. frequencies of occurrences, ranks, qualitatitive, or
mixed qualitative/quantititive data types).
- In correspondence analysis
such data (both rows/objects and columns/variables)
is mapped into a constant and identically weighted coordinate system.
- Note 1 on other agglomerative criteria:
It is a
very easy matter to allow for any of the other Lance-Williams criteria.
Note in particular that the computational time of the complete link method
using this NN-chain algorithm is O(n2). (Quite a number of
published papers erroneously claim a higher computational requirement.)
- Note 2 on timing experiments with my hclust program available
in the official
R distribution: this program additionally finds cluster assignments,
and that part of the processing is O(n3), which dominates.
- Note 3: for background see F. Murtagh, Multidimensional Clustering
Algorithms, Physica-Verlag, 1985. Scanned version of this book
available (see below).
2. MDA-J: Multivariate Data Analysis - Java
- A new area has been set up for this code, which has its own address: see
www.correspondances.info.
- For background see F. Murtagh, Correspondence Analysis and Data Coding
with R and Java, Chapman & Hall/CRC Press, 2005.
3. Multivariate Data Analysis Software as Individual Java Applications
- [Doc:]
Documentation on all programs which follow.
- [Sample data:]
Fisher iris data, iris.dat.
Format:
row and column numbers (integer), followed by sequence of
matrix data values (floating), read row-wise. The output obtained is
available in each case below.
- [PCA:]
Principal Components Analysis on correlations,
PCAcorr.java,
which requires the JAMA class library. (See below for
a link to JAMA.)
To run:
(i) Assume JAMA class library is in subdirectory Jama in
current directory.
(ii) Compile: javac PCAcorr.java
General note: if your classpath variable is not set, you can run
"javac" or "java" as: "javac -classpath ." and "java -classpath .".
(iii) Get syntax to standard output: java PCAcorr
(iv) Run on iris.dat: java PCAcorr iris.dat >
pcaoutput.txt.
-
Hierarchical clustering - background reading:
A small text, but one with everything necessary on the most effective
hierarchical clustering algorithms (i.e., using nearest neighbor chains,
and/or reciprocal or mutual nearest neighbors) is F. Murtagh,
Multidimensional Clustering Algorithms, Physica-Verlag, 1985. See
in particular the table on page 68.
[HCL:] Hierachical Clustering, HCL.java.
No additional class
libraries required. Currently the minimum variances criterion (Ward's
method) is supported. Changes for other
criteria (single, complete link, etc.) are indicated in the code.
To run:
(i) Compile: javac HCL.java
(ii) Get syntax to standard output: java HCL
(iii) Run on iris.dat: java HCL iris.dat >
hcloutput.txt.
- Correspondence analysis - background reading:
For theoretical background and thoroughly
comprehensive discussion of applications and practice, and most of all
providing a deep discussion of algorithms, see J.-P. Benzécri,
Correspondence Analysis Handbook, translated by T.K. Gopolan,
Marcel Dekker, 1992.
[CA:]
Correspondence Analysis on fuzzy coded input,
CAfuzzy.java,
which requires the JAMA class library (see below for a link).
To run:
(i) Assume JAMA class library is in subdirectory Jama in
current directory.
(ii) Compile: javac CAfuzzy.java
(iii) Get syntax to standard output: java CAfuzzy
(iv) Run on iris.dat: java CAfuzzy iris.dat >
caoutput.txt.
- Partitioning and k-means clustering - background reading: H. Späth,
Cluster Dissection and Analysis: Theory, Fortran Programs, Examples,
Ellis Horwood, 1985, covers various algorithms including the
exchange algorithm. (This algorithm is related to work by Simon
Régnier, at the Maison des Sciences de l'Homme around 1964.)
[Partition:]
Partitioning or k-means using minimal
distance criterion and exchange algorithm, partition.java,
which requires the JAMA class library (see below for a link),
because the initial clusters
are defined from principal component projections.
To run: i) Assume JAMA class library is in subdirectory Jama in
current directory.
(ii) Compile: javac partition.java
(iii) Get syntax to standard output: java partition
(iv) Run on iris.dat: java partition iris0.dat >
partoutput.txt.
4. Gaussian Mixture Modeling with Bayes Factors
This is a new area, where we will get - soon - programs in C uploaded,
mainly for image segmentation (including multiband images) based on
Markov random field models, and with use of Bayes factor inference
- Bayes information criterion and BIC in the pseudolikelihood case.
- Gaussian mixture model of a univariate data set, and BIC to
assess goodness of model fit. Program gmm-bic.c.
Sample input data oil.dat, oil prices from Ross.
To run: gmm-bic 3 749 oil.dat (3 = number of clusters, 749 =
number of observations or values in the input data file.) Output to
standard output device.
Output from the above.
(Reference for the Ross data: Sheldon M. Ross, An Elementary
Introduction to Mathematical Finance : Options and other Topics,
Cambridge University Press, 2002.)
- Version of the same program, but with the following normalizion used
by Ross: values analyzed are
log value(i+1) - log value(i). Program (only
difference: use of this preprocessing of the data).
Output from the foregoing.
- Coming soon: code for 2D and 3D single band, and 2D and 3D
multiband, image segmentation, using BIC or pseudo-likelihood information
criterion. (Used for work on multiband astronomy, Earth observation,
medical imagery - see publications list and recent papers accessible from
homepage.)
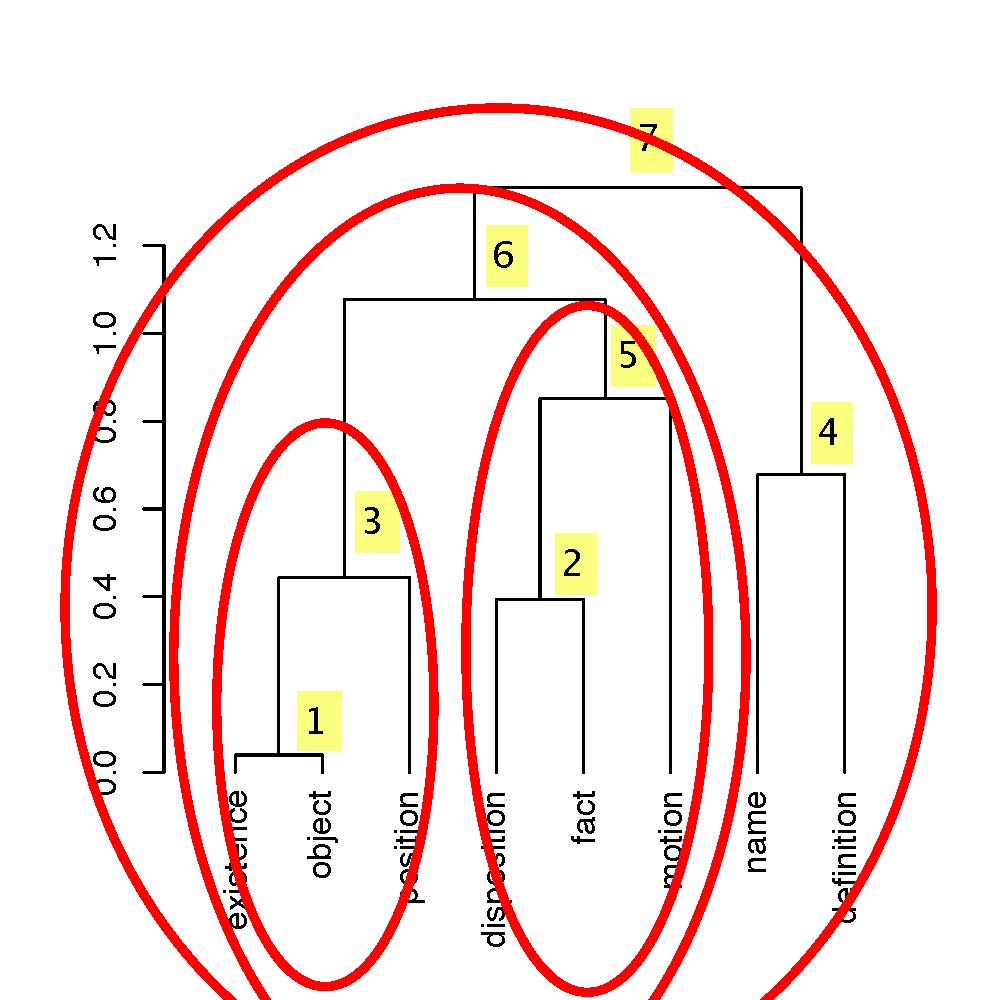
5. Wavelet Transform on a Hierarchy or Dendrogram
New hierarchical Haar wavelet transform
in R (see commented lines at start for example of use), which works on
a hierarchy produced by the foregoing hierarchical clustering programs.
This hierarchical Haar wavelet transform carries out the following
processing tasks: (i) from the data and a hierarchy, produce the
wavelet transform; (ii) filter the wavelet coefficients, using a
user-specified hard threshold; and (iii) reconstruct the data, i.e. perform
the inverse wavelet transform.
6. Multivariate Data Analysis Software in Fortran (and C)
The following is provided in case it is still of
interest. (Of note:
the agglomeration sequence visualization program.)
This is a collection of stand-alone routines, in Fortran (mostly) and C.
Sample data sets are available. Indications are given on how to compile, link
and run. Download the programs and run on your system. Many of these
programs were originally used on a VAX/VMS system, and later on Linux and
Solaris systems.
Please notify the author of any problems (although the programs are
provided "as is" and there are evident improvements which could be made).
The example of compiling, linking and running
given for principal components analysis (in Fortran) is similar to what is
required for the other Fortran programs here.
- S-Plus/R area. This subdirectory took all code
into a package. Listed below are standalone, individual programs.
- Principal components analysis (Fortran)
pcat.f, driver program
pca.f, routines used
spectr.dat, small sample dataset, originally
related to stellar spectra.
To compile and link: f77 pcat.f pca.f -o pcat
To run: pcat (output to screen, which may be directed to a file).
- Principal components analysis (C)
pca.c, program
To compile and link: cc pca.c -lm -o pcac
To run: pcac spectr.dat 36 8 R
Or: pcac spectr.dat 36 8 r
- Partitioning (Fortran)
partt.f, driver program
part.f, routines used
To compile and link: f77 partt.f part.f -o partt
Set up to run on spectr.dat (hard-wired in partt.f).
- Hierarchical clustering using stored dissimilarities (Fortran)
hct.f, driver program
hc.f, routines
hcass.f, cluster assignments routine
hcden.f, draw part of dendrogram
To compile and link: f77 hct.f hc.f hcass.f hcden.f -o hct
To run on spectr.dat (hard-wired): hct
- Hierarchical clustering without storage of dissimilarities (Fortran)
hcon2t.f, driver program
hcon2.f, routines
To compile and link: f77 hcon2t.f hcon2.f hcass.f hcden.f -o hcon2t
To run on spectr.dat (hard-wired): hcon2t
- Linear discrimant analysis (Fortran)
ldat.f, driver program
lda.f, routine
spectr2.dat, input dataset
- Multiple distriminant analysis (Fortran)
mdalum.f, driver program
mda.f, routine
luminosity.dat, input dataset
Additional diagnostics provided for this 3-class
supergiant - giant - dwarf dataset.
- K-nearest neighbors discriminant analysis - 2-class case (Fortran)
knn.f, driver program and routines used
ngc.dat, data - class number, followed by11
variables, for 500 objects.
The data is a set of 500 objects derived from an HST WF/PC image, in
the context of a study in star/cosmic ray hit discrimination.
To use: copy ngc.dat to ngc2.dat, to provide a second dataset. We
check assignments among the latter, relative to the assignments of objects
in ngc.dat. Hence, evidently for k=1 we must get exactly the correct
assignments in all cases. But for k=2 and higher, we may find
inconsistencies, which point to incorrect original class assignments.
Note: for k = even number, we allow for no assignment decision. I.e.
no clear majority.
Both idential or proportional prior situations are considered (i.e.
taking into account the relative numbers of class 1 and 2 memberships,
or not).
Also we run through all cases from k = 1, 2, ..., 15, which takes some
time.
To run: f77 knn.f -o knn
Then: knn (which takes ngc.dat and ngc2.dat as inputs, and produces
output on screen).
- Correspondence analysis (Fortran)
cat.f, driver program
ca.f, routines
uno.dat, input data (UNO voting)
- Classical or metric multidimensional scaling (Fortran)
cmdst.f, driver program
cmds.f, routines
cities.dat, input dataset
Currently this program generates its own random input data.
- Sammon mapping (Fortran)
sammon.f, program
iris.dat, input dataset (Fisher's iris data)
- Kohonen self-organizing feature map (Fortran)
koh.f, program
iris_norm.dat, input dataset (normalized
Fisher's iris data)
iris.f, for information, program to normalize data
- Sort routine (Fortran)
sort.f
- Weighted Levenshtein or edit distance between character strings (Fortran)
levenshtein.f
- Errors-in-variables regression, 2-dimensional (Fortran)
leiv1.f, York (Can. J. Phys. 44, 1079-1086, 1966)
leiv2.f, Fasano and Vio
leiv3.f, Ripley
All these programs are set up with sample driver routines and sample data.
They should perform exactly the same task.
- Minimal spanning tree, efficient algorithm in 2 dimensions (Fortran)
mst2d.f, program due to F.J. Rohlf, SUNY
- GMDH - group method for data handling, or Ivakhnenko polynomial
method (Fortran)
gmdh.f, program
gmhd.dat, sample data
See book by Farrow on GMDH.
- Point pattern matching approaches, software, data sets.
Preliminary web
page.
7. Links and Resources
- F. Murtagh, Multidimensional Clustering Algorithms,
Physica-Verlag, 1985.
Title pages and preface,
Contents,
Chapter 1,
Chapter 2,
Chapter 3,
Chapter 4,
Chapter 5.
- F. Murtagh
(with A. Heck), Multivariate Data Analysis, Kluwer, Dordrecht, 1987.
An expanded version of this
work is available online as a
271-page PDF file.
- Lectures, presentations on principal components
analysis, correspondence analysis,
other dimensionality reduction methods, discriminant analysis,
cluster analysis, with various applications.
- Collection of time series,
used in, among other work, F. Murtagh, "Identifying the
ultrametricity of time series", European Physical Journal B, 43, 573-579, 2005.
- [Data sets:]
Time
series data library. Over 800 time series from various domains. Very
useful.
- Prof. J.P. Benzécri's original
correspondence analysis code in Pascal, for Macintosh OS 9 (or earlier)
platform.
- NewMDSX - Multidimensional
Scaling, including CANDECOMP, HICLUS, INDSCAL-S, MDPREF, MINI-RSA,
MINISSA, MRSCAL, PINDIS, PREFMAP, PRO-FIT, TRISOSCAL, WOMBATS. Windows
executables and documentation.
For original programs, see
Bell Labs MDS programs.
- Visual user interfaces, using Kohonen
self-organizing feature maps and other data display approaches.
- FITS Table and Image File Reading in R: Currently only reading
FITS (Flexible Image Transport System)
Table or Image files in Windows is supported.
File rfitsio.zip, containing two dll files to
be loaded, and four R files to get height, width and total size information,
and to read the FITS file. Uses CFITSIO libraries from GSFC.
- Pointers
to, and addresses of, lots of
multivariate data analysis code, a text file collected or summarized
by F. Murtagh from mail messages or digest announcements. Look here for
whereabouts of code for: decision trees, clustering (C code),
multidimensional scaling and lots of other psychometric mapping methods,
Voronoi diagrams, etc.
- Statlib,
major repository of
statistical software, datasets, and information such as email lists and
organisational addresses, at Carnegie Mellon University (Mike Meyer,
[email protected]).
- Weka Machine Learning
Project, Waikato Environment for Knowledge Analysis,
machine learning workbench.
- Mixture modeling page.
- Java resources.
- [Multiresolution analysis:]
Examples of application. Matlab (and IDL)
software,
accompanying the book Sparse Image and Signal Processing:
Wavelets, Curvelets, Morphological Diversity, JL Starck, F Murtagh
and J Fadili, Cambridge University Press, 2010. Some other sites
with software, a large
package, executables for common platforms, and
for Windows PCs.
Author:
fmurtagh at acm dot org
Homepage